Millsaps College
Become Something Major
Become Majorly You
Are you looking for more from college than just being equipped for a career? Do you want a place that helps you become the best version of yourself? A place that offers academic excellence, hands-on learning and leadership opportunities in the classroom and in the community?
We believe college shouldn’t be a transaction, it should be transformational! Come experience the major opportunities at Millsaps.
Major Possibilities
Creative Thinkers.
Solution Finders.
Future Changemakers.
Become part of a community that seeks to challenge perspectives, create beauty and effect positive change in the world. Our experiential learning opportunities allow you to apply your knowledge and skills in real-world settings. These experiences equip you to move into career leadership roles faster.
9:1
9-to-1 student-to-teacher ratio
18
Number of NCAA Division III sports
1st
1st Phi Beta Kappa chapter in Mississippi

Learning and Community
The Major Difference
Academics
Opportunities for transformative learning and leadership experience.
Campus Life
Find yourself and create lasting connections.
Pathways
Six specialized tracks to help you develop and explore.
The Millsaps Family
“When I first stepped foot on campus, I knew then I would find my place and myself here.”
News & Events
MAJOR Happenings

Latest News
Two Millsaps Students Earn National ODK Leadership Honors
Students from across the country are considered, and selection reflects not only exceptional achievement, but leadership that positively influences campus culture and the broader community.

Millsaps Hosts Forum on the American Experiment
These conversations reflect the kind of engagement we’re building at Millsaps—where students learn to navigate complex issues with thoughtfulness and respect.
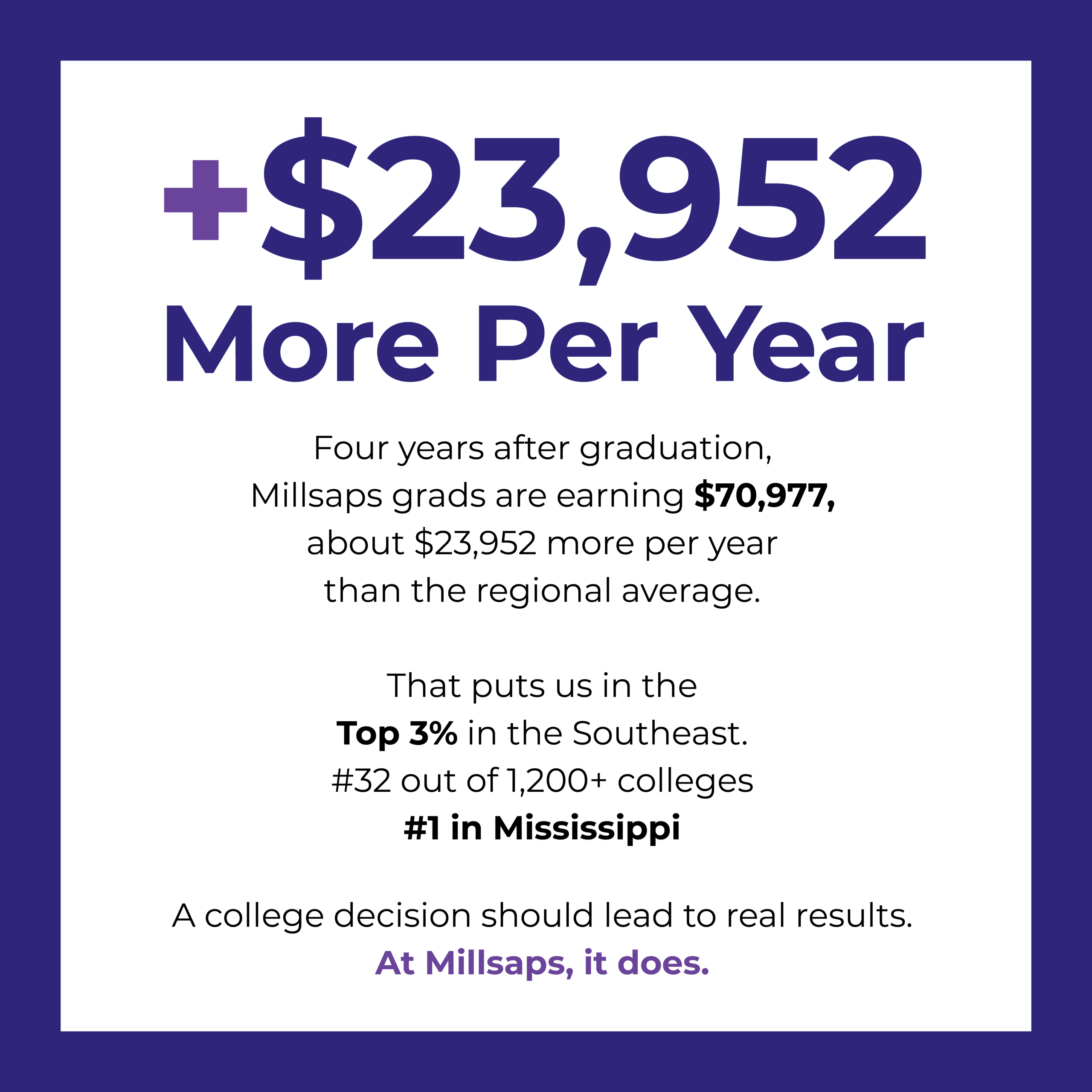
Millsaps Grads Earn More, New Federal Data Shows
The post-graduation achievements of Millsaps alumni reflect our commitment to the success of every student.

Reverend Carr to Be New Dean of the Chapel at Millsaps
Millsaps has certainly helped shape who I am, and I know first-hand what a special and transformative place it is.

